What are protein interaction networks?
Protein interaction networks provide information about how proteins interact to form complexes that can have specific functions [1]. When one protein in a network is modified, many interactions can potentially be disrupted and certain cell functions or processes can be affected.
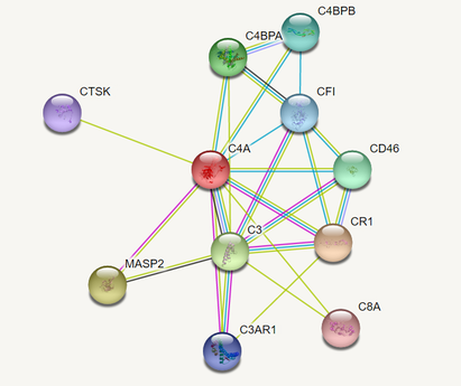
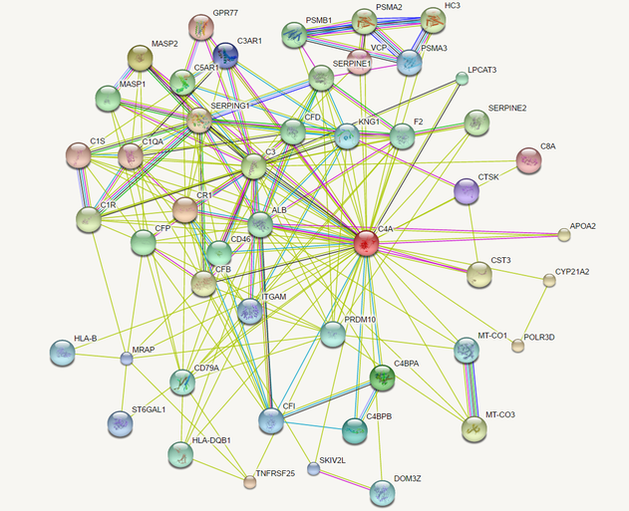
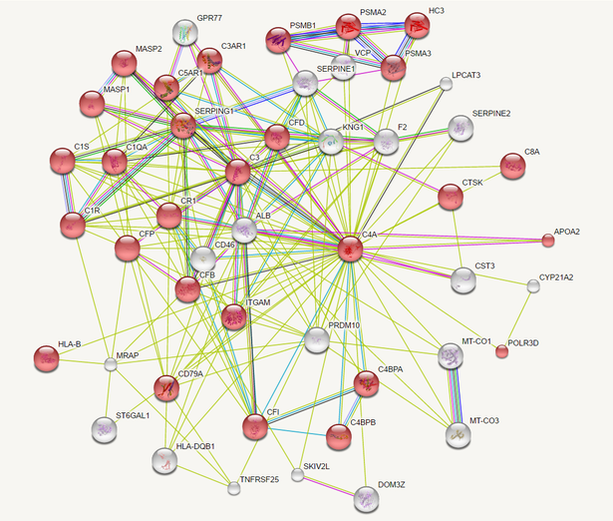
C4A Interaction Networks
There are several ways to generate a map of protein interactions, however I used the STRING database. STRING is a versatile database that reveals predicted as well as known protein-protein interactions, and also allows the user to change various settings and to access the primary literature that described each interaction.
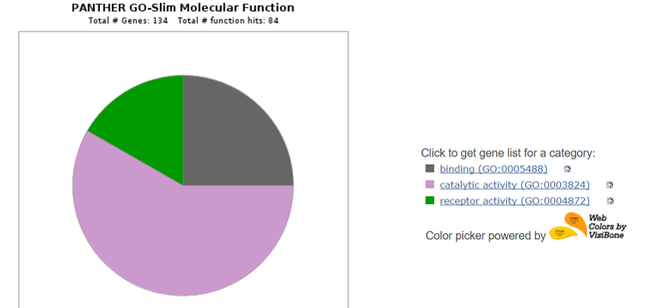
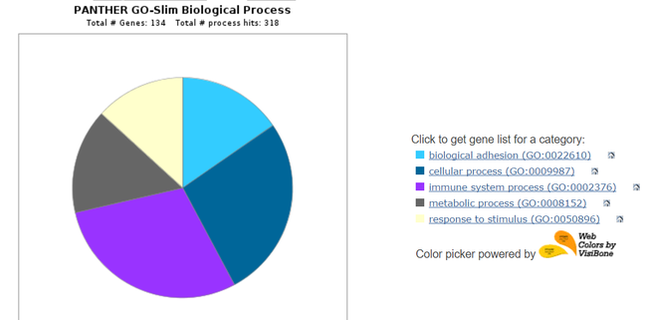
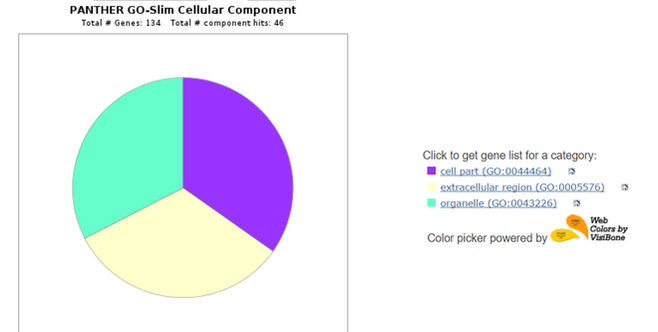
After downloading the list of interacting proteins on STRING, the list can be uploaded onto the PANTHER Classification System. PANTHER can compare the gene ontology terms for each interacting protein and display common classifications in a pie chart. This is helpful for observing what functions the interacting proteins have in relation to C4A, as well as for determining novel pathways or functions that could be of interest. Below are three pie charts obtained from a PANTHER search of human C4A interactors.
Discussion
The human C4A protein interacts with several other proteins directly and with many proteins through more distant interactions. Many of the interacting proteins are other complement components, and 27 of the 47 interacting proteins are involved in regulation of immune response. A significant number of the interactors also had "immune system process" as one of their biological process GO terms. Studying these interactions can tell researchers more about how C4A works while helping to reveal the mechanism of synaptic pruning.